Using the human brain to understand “omics” in autism
by Shannon Ellis, Ph.D. at Johns Hopkins University

Autism is a complex neurodevelopmental disorder with a definitively established genetic basis1. However, complete understanding of the genetic etiology remains elusive. While the role for certain CNVs in autism has been definitively established2,3 and exome sequencing studies have begun to uncover rare de novo mutations that play a role in the disorder4–9, we are far from identifying the names of the hundreds of genes likely contributing to the disorder.
While naming the genes that play a role in autism is critical, it has become increasingly clear that changes in the DNA sequence are only a first step toward complete understanding. Additionally, scientists acknowledge that they must know what the genes do, not only what they are. Therefore, our lab has begun to get a handle on what is altered at the level of gene expression and DNA methylation within the primary affected tissue in autism – the brain. This is part of a larger field called “omics”: meaning “genomics” (the study of DNA sequence), “transcriptomics” (the study of gene expression), “epigenomics” (the study of how genes are expressed, “proteomics” (how proteins are regulated by DNA expression) and “metabolomics” (the chemical processes of making and breaking down compounds). Our study focused on transcriptomics and epigenomics.
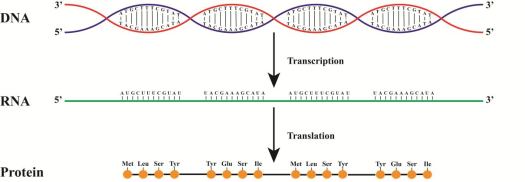
To understand the processes in the brains of people with autism, we had to study the brains of people with autism. Therefore, we worked with the Autism BrainNet (formerly the Autism Tissue Program) to obtain brain samples and extract RNA. We were interested in RNA because it is produced from DNA. Using DNA as a template, RNA is formed as the first step in creating proteins. The proteins are what carry out the function of the cell. The number of copies of RNA is a reflection of the gene expression of the cell: the more copies, the more gene expression. While differences in an individual’s DNA are undoubtedly of interest to the study of autism, it is also important to look for differences in gene expression and protein levels to fully understand the disorder. We used the RNA to look at the gene expression of about 14,000 genes. Additionally, in these same individuals, DNA samples were used to estimate methylation levels at cytosines across cytosine-rich regions of the genome.
Last year, we published some results on the transcriptomic part. When we looked at the gene expression, we identified three groups of genes that showed different patterns of expression between autism cases and controls. The first two groups included genes that influence the way neurons work, how they interact with each other, and how they grow and communicate to form the human brain. Interestingly, the genes that showed differences in gene expression were different genes than those identified in previous studies that, rather than looking at gene expression differences, looked to identify DNA differences important to autism. This demonstrates that genes that have differences in their DNA are different from those genes showing downstream differences at the level of gene expression. The third group was made up primarily of M2-microglia genes, suggesting in increased immune response in the brains of autistic individuals. We want to stipulate that this does not mean that alterations in the immune system of the brain cause autism. It could be that abnormal gene expression in the brain triggers an M2 microglia response. Future work is required before we can determine if the increased immune response leads to or is a result of autism. Nevertheless, from a treatment standpoint, this work provides pathways that, despite variable genetic causes, can be targeted for treatment in affected individuals going forward.
We moved this further to compare the gene expression overlap in the brains of people with autism with other neuropsychiatric disorders. Previous studies have shown that there is an overlap in which genes have DNA differences across neuropsychiatric disorders. To establish if this overlap holds up at the level of the transcriptome, we compared gene expression differences across three disorders: autism, schizophrenia, and bipolar disorder. While there was little overlap between autism and bipolar disorder, there was significant overlap between the expression patterns in genes in autism and schizophrenia brains11, with consistent decreased expression at neuronal and synaptic plasticity genes across these two disorders. This work extended the known genetic overlap between neuropsychiatric disorders by establishing a relationship between alterations in gene expression found in both autism and schizophrenia.
Finally, as gene expression is directly regulated by DNA methylation, we looked to determine if DNA methylation differences play a role in autism. Methylation is a process where a methyl group attaches to a part of a DNA sequence and turns down the expression of that gene. To look at DNA methylation, we looked at cytosines. Cytosines are places in our genome where these methyl groups normally attach. Thus far, most work has studied CpG methylation. This refers to when a methyl group attaches to a cytosine (C) that is directly next to a guanine (G) nucleotide. This CpG context is where methylation most frequently occurs in the genome. However, methylation can occur at cytosines next to other DNA nucleotides (C, T, or A), and this is referred to as CpH methylation. As CpH methylation occurs at higher levels in the brain relative to other tissues, we did not want to limit our study to CpG sites alone, but rather wanted to look for differences in CpH methylation as well. We found increased levels of methylation at CpH, but not CpG, sites globally within the autistic brain. We are currently working on why there is this difference and why it is seen in autism, but as CpH methylation is largely specific to the human brain (as compared to blood or other cells), it is a particularly compelling finding.
While we acknowledge we have not answered all the questions, we now have a better understanding of what is going on in the brain of individuals with autism. In particular, in addition to further establishing a relationship between schizophrenia and autism, this work not only highlights a role for increased immune activation within affected individuals, providing a particular pathway to target when considering future therapies, but also, for the first time, suggests a role for increased methylation at cytosines within the CpH context in the brains of autistic individuals.
References:
- Gaugler, T. et al. Most genetic risk for autism resides with common variation. Nat. Genet. 46, 881–885 (2014).
- Marshall, C. R. & Scherer, S. W. Detection and characterization of copy number variation in autism spectrum disorder. Methods Mol. Biol. Clifton NJ 838, 115–135 (2012).
- Sebat, J. et al. Strong Association of De Novo Copy Number Mutations with Autism. Science 316, 445–449 (2007).
- Sanders, S. J. et al. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature (2012).
- Yu, T. W. et al. Using Whole-Exome Sequencing to Identify Inherited Causes of Autism. Neuron 77, 259–273 (2013).
- Iossifov, I. et al. De Novo Gene Disruptions in Children on the Autistic Spectrum. Neuron 74, 285–299 (2012).
- O’Roak, B. J. et al. Exome sequencing in sporadic autism spectrum disorders identifies severe de novo mutations. Nat. Genet. 43, 585–589 (2011).
- Neale, B. M. et al. Patterns and rates of exonic de novo mutations in autism spectrum disorders. Nature (2012).
- De Rubeis, S. et al. Synaptic, transcriptional and chromatin genes disrupted in autism. Nature 515, 209–215 (2014).
- Gupta, S. et al. Transcriptome analysis reveals dysregulation of innate immune response genes and neuronal activity-dependent genes in autism. Nat. Commun. 5, 5748 (2014).
- Ellis, S. E., Panitch, R., West, A. & Arking, D. E. Transcriptome Analysis of Cortical Tissue Reveals Shared Sets of Down-Regulated Genes in Autism and Schizophrenia. bioRxiv 29132 (2016).
